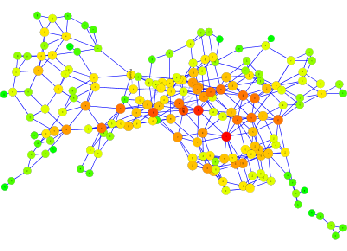
Part of the network of chemical reactions for about 150 biochemical reactants with <=3 carbon atoms.
 The bacterium E. coli which is a common cause of urinary tract infections is thought
to live primarily in the large intestine.
The bacterium E. coli which is a common cause of urinary tract infections is thought
to live primarily in the large intestine.
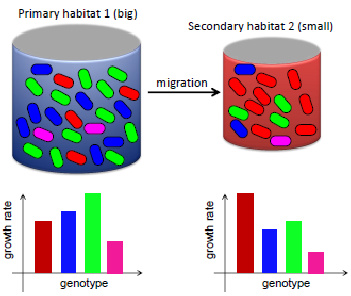
In our model, bacteria migrate from a large, primary habitat to a smaller, secondary habitat where different genotypes have different growth rates compared to the “source” habitat.

Evolution of drug resistance takes a completely different course for a uniformly
distributed drug (left) than for a drug gradient (right). Uniform drug distribution
causes the black strain (susceptible) to evolve into the blue one (resistant) at
random places in the environment. This strain may then evolve to a more resistant
one (green) etc. In contrast, drug gradient cause the population to evolve in waves
of increasingly better adapted mutants and to extend its range in a step-
Evolution of drug resistance
Drug resistance is a very important problem in the treatment of bacterial or viral (such as HIV) infections, and in cancer. For example, many bacteria have become resistant to antibiotics such as penicillin, and some bacteria (“superbugs”) are simultaneously resistant to many antibiotics, which makes it very difficult to kill them.
The reason is that pathogenic organisms have a natural ability to evolve and adapt
to harsh conditions caused by therapy. Such micro-
Other areas of interest
-
-
-
Although majority of mutations is deleterious, some of them make cells more resistant to drugs. Frequent application of any drug causes the population of pathogenic cells to become enriched in resistant strains, because susceptible pathogens are killed whereas resistant ones thrive. Therefore, any drug becomes inefficient after some time. An important question is how long does it take?
I investigate how bacteria become resistant to antibiotics. In particular, I study how spatial inhomogeneities such as drug gradients affect the rate of adaptation of
bacteria. I have recently published (with P. Greulich and R. Allen) a theoretical paper about this problem and I am currently making some experiments to confirm our theory.
Simple models for bacterial evolution with migration
In nature, evolution proceeds not only through mutation and selection of better adapted
organisms within a uniform population, but also through migration of organisms between
different environments. The role of migration in asexual microbes was long underestimated.
However, recent experiments have suggested that many pathogenic bacteria which cause
infections are not in their native environment but are actually colonisers from nearby
environments where they are non-
In my research, I investigate the role of migration in the evolution of genetic diversity in microbes. I have published (with R. Allen and M. Evans) a paper in which we show that moderate migration strongly increases genetic diversity, but too strong migration inhibits adaptation to the new environment. I am also working on how this scenario is modified in the presence of cooperation between different bacterial species, horizontal gene transder, and predation by phages (viruses that infect bacteria).
Metabolic pathways and microbial diversity
Some metabolic pathways (e.g. glycolysis, the pathway which metabolises the sugar glucose) are almost identical in all organisms. We (me, S. Court, R. Allen, and P. Warren) are interested in how these pathways might have evolved. In particular, we try to understand why biological evolution has selected specific chemical reactants and chemical reactions in those pathways. We have generated a network of feasible, enzymatically catalysed chemical reactions for organic molecules with up to 6 carbon atoms and are investigating alternative pathways to glycolysis in hope to understand what makes the real pathway so universal.
Growing bacterial colonies
I use computer simulations (with F. Farrel, D. Marenduzzo, and O. Hallatschek) to
study the growth and ordering of rod-
I work with R. Allen and Pietro Cicuta on the transition from 2d to 3d growth in bacterial colonies trapped between a glass surface and an elastic agar slab.
I also work (with F. Farrel and O. Hallatschek) on on the establishment rate of new mutations in bacterial populations.
My PhD student Kuba Pastuszak simulates conjugation (a form of horizontal gene transfer) in bacterial colonies.
Rod-
Biophysics
Spatial models of cancer
I work with M.A. Nowak, I. Bozic, and B. Vogelstein on spatial models of cancer progression. Cancer occurs when enough mutations accumulate to cause uncontrolled cell growth. Along with “driver” mutations that affect the rate of growth/death, many “passenger” mutations occur which initially do not confer any fitness advantage but may do so when conditions change, for example, as a result of chemotherapy. I am interested in how growth in space affects genetic diversity of solid tumours, the number of such passenger mutations, and the probability of tumour regrowth.
My PhD student Chay Paterson works on computer/mathematical models of tumors composed of multiple lesions (see the video below).
Example of a simulated solid tumour. Similar colours correspond to genetically similar cells. A half of the tumour has been cut off to better visualise its interior. You can see that different parts of the tumour have accumulated different mutations.
Simulation of a tumour composed of multiple lesions. When treatment begins, most
of the tumour dies. This makes space for resistant cells that have occurred through
spontaneous mutations during tumour growth and causes the tumour to re-